Reporting on the #SciWri16 #3Dgenome session.
If you have taken any genetics courses in high school or college, this top image might be familiar to you. Inside a cell, there is the nucleus, which houses the chromosome, and the chromosome is made of the DNA.
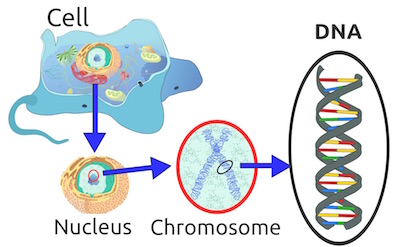 DNA is an astounding molecule. If we were to stretch out a DNA molecule, it would be about 2 meters long (around 6.5 feet). The nucleus that houses the DNA is only 6 micrometers in size, which is about 100 times smaller than a grain of salt. That is like trying to stuff a thread that stretches from Philadelphia to Washington, D.C. (140 miles) in a check-in luggage bag (27 inches in width). And, to make this even more complex, the DNA cannot simply get haphazardly packed into the nucleus. It has to remain functional so the cell can produce functional proteins and survive — meaning that how it is packed matters.
DNA is an astounding molecule. If we were to stretch out a DNA molecule, it would be about 2 meters long (around 6.5 feet). The nucleus that houses the DNA is only 6 micrometers in size, which is about 100 times smaller than a grain of salt. That is like trying to stuff a thread that stretches from Philadelphia to Washington, D.C. (140 miles) in a check-in luggage bag (27 inches in width). And, to make this even more complex, the DNA cannot simply get haphazardly packed into the nucleus. It has to remain functional so the cell can produce functional proteins and survive — meaning that how it is packed matters.
So how does the DNA fit into the nucleus? To answer that question, Baylor College of Medicine geneticist Erez Lieberman Aiden and his team are investigating how genome remains functional in spite of being packed into such tight space.
“Usually when we are packing things in backpacks or car trunks, I can take it all out later,” Lieberman Aiden said. “That’s not case for the genome. The genome has to stay in the nucleus forever. The cell doesn’t ever have space to expand. The problem of how you package a genome into the cell has layers of complexity that most packing problems we encounter don’t have. Most packing problems you encounter are about packing in minimized space. The genome has to be packed in minimized space and stay incredibly functional.”
Contact Map Reveals 3D Genome Structure
At the ScienceWriters2016 conference, Lieberman Aiden gave a talk called The Human Genome’s 3D Code to explain the way he and his colleagues study how DNA is organized and packed in the nucleus. See below for a Storify archive of this #3Dgenome session’s tweets [Roca 2016].
Lieberman Aiden described a tool that he helped develop as a graduate student called Hi-C — a play on the names of similar research tools (3C, 4C, and 5C) and the name of a popular juice [Lieberman-Aiden 2009]. The tool is designed to reveal which parts of the genome are close to each other.
 Using Hi-C, Lieberman Aiden and his colleagues were able to develop a “contact map,” which shows the physical distance between the two regions of the genome. To illustrate how Hi-C works, Lieberman Aiden made a contact map for characters from the television show The Simpsons. The contact map shows how many times two characters are in a picture together, which indicates how close the two characters are. In the example, Homer has many more pictures with Marge than with other characters, suggesting Homer and Marge have a close relationship and interact a lot.
Using Hi-C, Lieberman Aiden and his colleagues were able to develop a “contact map,” which shows the physical distance between the two regions of the genome. To illustrate how Hi-C works, Lieberman Aiden made a contact map for characters from the television show The Simpsons. The contact map shows how many times two characters are in a picture together, which indicates how close the two characters are. In the example, Homer has many more pictures with Marge than with other characters, suggesting Homer and Marge have a close relationship and interact a lot.
The same goes for two regions of the genome that show a close physical distance in the Hi-C contact map. The closer the two regions are, the more they interact. The two regions might be close to each other and interact often because they have a functional relationship, like Homer and Marge. And based on the contact map, researchers can get a glimpse into how the genome is packed into the tiny nucleus.
3D Structure Shows Link between Structure and Function
After analyzing the contact map, the researchers came to two main conclusions. First, all parts of the human genome roughly falls into one of two compartments that are physically located in different places in the nucleus. Lieberman Aiden described the compartments as “social clubs,” in which a particular regions of the DNA can “hang out” with other DNA sequences in either club; but, it is more likely to hang out in one more than the other. One compartment is comprised of the “active” genome, which expresses genes to produce proteins. The other compartment is the “inactive” genome, where genes are not expressed.
Based on these observations, it was clear to the researchers that the genome structure and its functions are closely tied. Because of this link, Lieberman Aiden said that the folding of the DNA is a dynamic process, in which the structural changes accompany functional changes. He compared the dynamic changes of the genome structure to a person’s re-folding of a newspaper, emphasizing that people fold newspapers differently depending on what stories they want to read.
“How do you read a newspaper? Newspapers actually have a lot of information; but, you can constantly change how you fold the information in order to access what you want. Usually when you are reading the newspaper, you don’t throw out the sections. You have all the sections. The question is how do I make the bins that I’m using accessible? The way you fold [the newspaper] has a huge impact on the prominence of different pieces of information.”
3D Structure Illustrates Our Genome’s Complexity
Human genomes consists of large loops, which Lieberman Aiden illustrated using blue cupcakes in his presentation. This conclusion is significant because it changes how we think about our genome. Human (and other mammalian) genomes are not linear and simple like in bacteria, where there are no complex 3D genome structures. Loops allow parts of the genome to influence distant parts of the genome — parts of the genome, like enhancers, that are important for a gene’s function no longer have to be right next to each other.
According to Lieberman Aiden, this conclusion from the Hi-C experiment illustrates how complex our genome is. This is a concept that still remains a curiosity for many researchers including himself. What could be the purpose of such complexity? Short answer — no one knows. But researchers are certain about one thing — the loops and other complex structures are clearly important. According to Lieberman Aiden, they are very well preserved across different mammalian species. If the loops are not important, there is no reason for different species to still have them.
Still, Lieberman Aiden said the human genome seems “inexplicably messy” compared to less-complex organisms like bacteria.
“We don’t really know why the human genome is designed this way,” Lieberman Aiden said. “There’s a joke — if you are on the committee of designing the smaller genomes, you would gladly take credit for it. But if someone accused you of being on the committee of designing the human genome, you would be embarrassed.”
Investigating Human Genome While Being One’s Strongest Advocate
Constructing the contact map and revealing the genome’s 3D structure would not be possible without tireless work from a team of scientists. Neva Durand, a staff scientist at Lieberman Aiden’s lab, helped to create two software programs for analyzing Hi-C data: Juicebox, which helps researchers visualize the data, and Juicer, which transforms raw Hi-C data into files that can be visualized using Juicebox.
 Durand said she initially began her studies in computer science; but, she decided to work with Lieberman Aiden in order to pursue interesting questions and prevent herself from feeling constrained in her previous field of study.
Durand said she initially began her studies in computer science; but, she decided to work with Lieberman Aiden in order to pursue interesting questions and prevent herself from feeling constrained in her previous field of study.
“Erez has lots of very cool ideas; and, he likes looking at a diverse array of problems in many different areas,” Durand said. “This aligns really well with me. I love learning new things and didn’t want to be stuck on one tiny little branch of research. It’s a very exciting, fast-moving field, and I learn more and more every day. The work is intense but a lot of fun.”
Having started her studies in computer science, Durand said she is familiar with feeling outnumbered as a female researcher. She said it is imperative for female and minority researchers to find a voice and be one’s own strongest advocate. Although some in technical disciplines might not appreciate these qualities in women, such advocacy is necessary for female career advancement.
“I think it’s always challenging and some people face more challenges than others,” Durand said. “And it’s completely unfair. However, given that the system is biased the way it is, it’s important to believe in yourself and advocate for yourself. I think especially members of minority groups have trouble believing in themselves, even when by any objective member they are succeeding.”
Durand also stressed the importance to find a supportive mentor who could provide the best environment possible for one to move forward.
“And it’s equally important to find good mentors that believe in you and will give you encouragement and that extra push it takes to succeed,” Durand said. “I’ve been very lucky to always find excellent mentors — both male and female — at every stage. They were very supportive at every stage of my career. This has continued with Erez, who is a fantastic mentor.”
I. Amato (2015) Genetic Geometry Takes Shape, Quanta Magazine, February 25
E. Lieberman-Aiden, et al (2009) Comprehensive Mapping of Long-Range Interactions Reveals Folding Principles of the Human Genome, Science, 326(5950):289-293
A.I. Roca (2016) #SciWri16 #3Dgenome, Storify, October 31
(top) Depiction of eukaryotic cell DNA. (Credit: Wikimedia)
(middle) Contact Map of Simpsons characters. (Credit: Ian Street)
(bottom) Neva Durand. (Credit: N. Durand)
I. Park (2016) Bringing DNA Structure and Function to the 3D Realm. DiverseScholar 7:5
Diverse Scholar is now publishing original written works. Submit article ideas by contacting us at info@DiverseScholar.org. This work is licensed under a Creative Commons Attribution-Noncommercial-No Derivative Works 3.0 Unported License.
![]()
Originally published 31-Dec-2016.
